The recent advent of chromatin conformation capture technologies has made it possible to systematically investigate spatial interactions between genomic loci at unprecedented resolution. This study provides a reference for the analysis of chromatin domains from Hi-C experiments and useful guidelines for choosing a suitable approach based on the experimental design, available data, and biological question of interest. Performances of these methods differ based on data resolution and normalization strategy, but a core set of TAD callers consistently retrieve reproducible domains, even at low sequencing depths, that are enriched for TAD-associated biological features. We find that TAD sizes and numbers vary significantly among callers and data resolutions, challenging the definition of an average TAD size, but strengthening the hypothesis that TADs are hierarchically organized domains, rather than disjoint structural elements. Here, we test and compare 22 computational methods to identify TADs across 20 different conditions. The reliability and reproducibility of these findings are intrinsically related with the correct identification of these domains from high-throughput chromatin conformation capture (Hi-C) experiments. 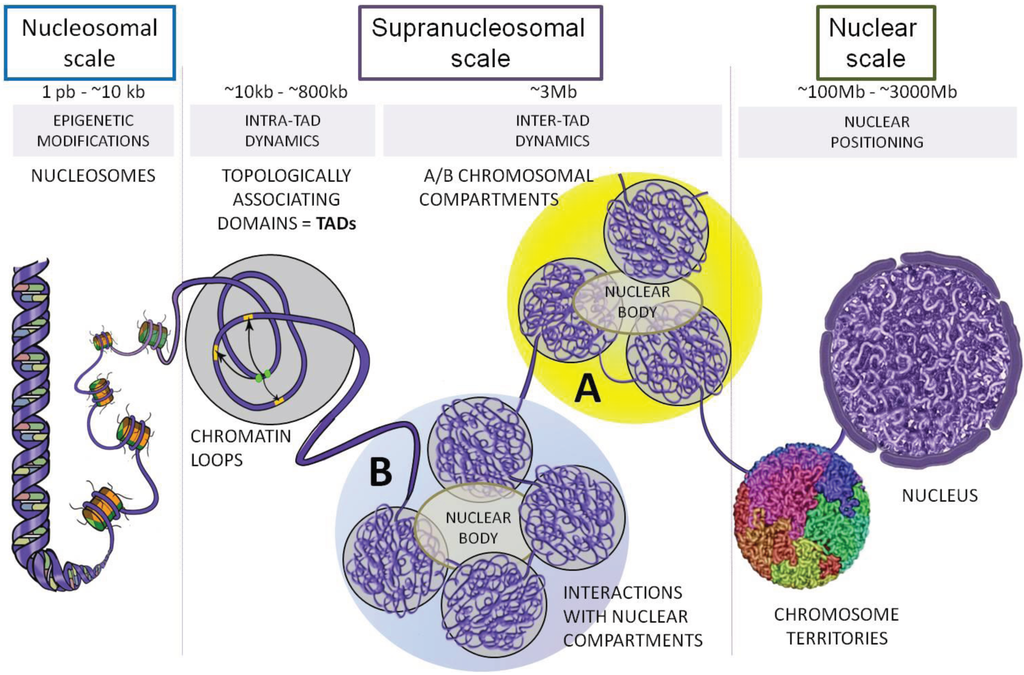
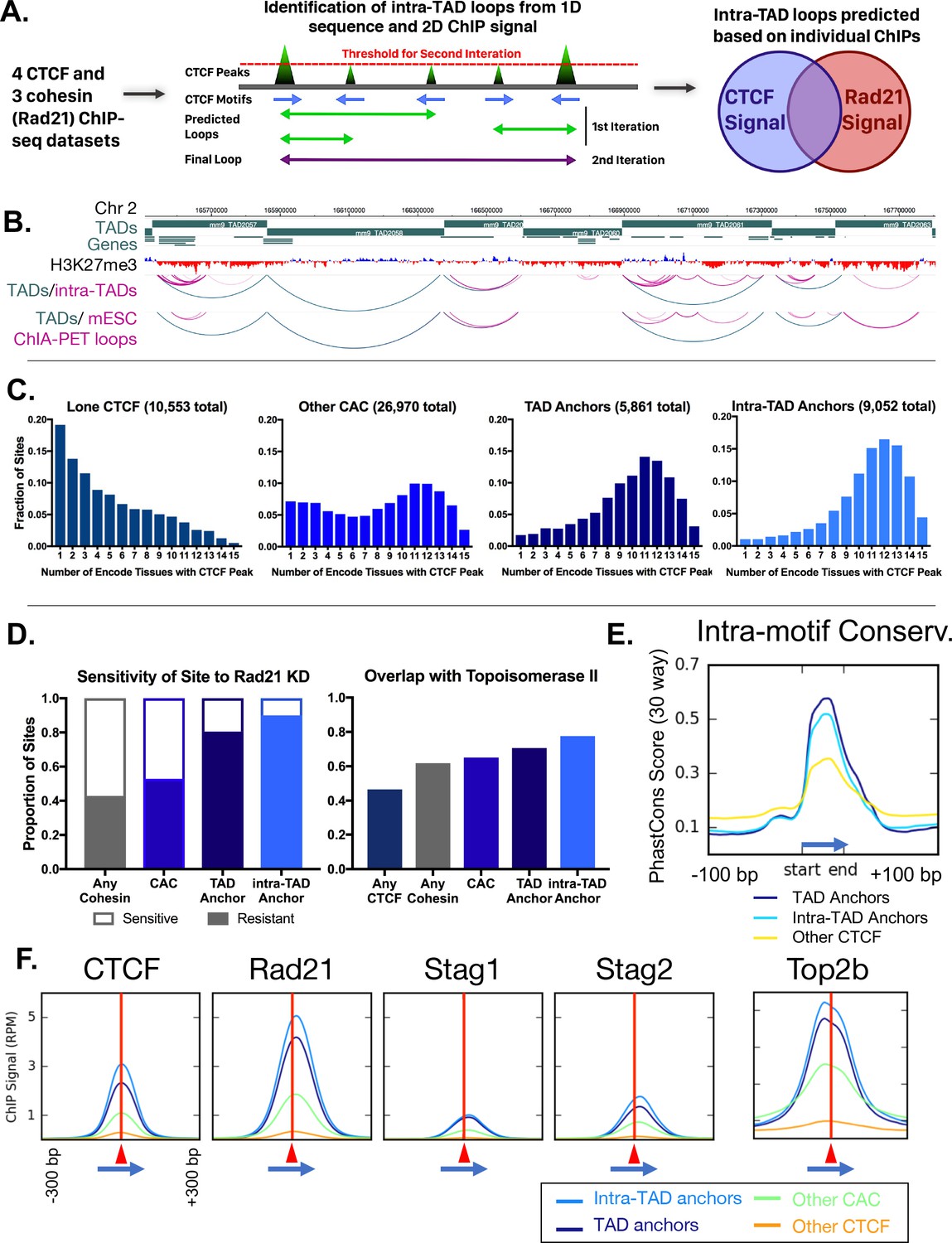
TADs have been characterized across multiple species, tissue types, and differentiation stages, sometimes in association with regulation of biological functions. Chromatin folding gives rise to structural elements among which are clusters of densely interacting DNA regions termed topologically associating domains (TADs).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
